Lujiong Danzhen, Bowen Jiang, Jack Markert, Ethan Tan, Guanlin Wu, Maizhe Yang ▫ Western Reserve Academy, Hudson OH
Reviewed on 6 May 2023; Accepted on 13 July 2023; Published on 16 October 2023
With help from the 2023 BioTreks Production Team.
Prolonged exposure to formaldehyde may cause severe eye and respiratory irritation and increase various cancer risks. Therefore, determining formaldehyde concentration in the air and regulating it to a safe level has become a priority for places with potential formaldehyde exposure risks. Current detection methods include using a mechanical detector, conducting lab samples, and sniff testing. However, these methods are often unreliable, expensive, and dangerous. We propose an economical and effective detector based on the ΔfrmR strain of Escherichia coli. The detector will be yellow by default and turns red when formaldehyde is present. We will construct the formaldehyde-detecting plasmid based on the R0010 (pLacI)_AB plasmid backbone and transform it into cells. Using a Pfrm promoter, the detector reacts to formaldehyde at levels around 100 μM. We then utilize the Plac-lacI repressor system as a genetic switch: with the presence of formaldehyde, lacI, a Plac repressor, is expressed to deactivate the yellow pigment and turn the cell red. The bacteria will be cultured in lactose-rich media to ensure the constitutive expression of the yellow pigment under Plac. Moreover, the frmA and frmB genes in the construct remediate formaldehyde in the solution. As the concentration of formaldehyde decreases through frmAB in the detector solution, the detector reverts to yellow, making the detector system reusable when provided with sufficient nutrients.
Keywords: Formaldehyde, formaldehyde detector, ΔfrmR strain, Escherichia coli, Pfrm, Plac-lacI
Authors are listed in alphabetical order. Beth Pethel from Western Reserve Academy, Hudson OH mentored the group. Please direct all correspondence to pethelb@wra.net.
Formaldehyde (CH2O) is a colorless poisonous flammable gas used extensively in preserving cadavers, insulating building materials, inactivating viruses in the production of vaccines, and many other industrial settings. Despite its wide applications, exposure to formaldehyde can cause severe eye, skin, and respiratory irritation. To combat this problem, our study aims to design an E. coli-based detection system to alert users of potentially harmful concentrations of formaldehyde.
Often used as a sterilant, formaldehyde kills microorganisms and is widely used to preserve organic materials such as biological specimens (National Institutes of Health, 2023). The antimicrobial and conservatory properties of formaldehyde come from its ability to generate protein linkages by binding to amino acids like lysine and tyrosine (Centers for Disease Control, 2016). This property also makes formaldehyde an ideal insulation and adhesive agent in building materials, such as particleboard and plywood (Environmental Protection Agency, 2022). In addition, numerous vaccines use small amounts of formaldehyde to inactivate the virus (e.g., polio vaccines) and detoxify bacterial toxins through the alkylation of proteins and purine bases (Common Ingredients, 2019; Herrara-Rodriguez et al., 2019).
In addition to its extensive industrial usage, formaldehyde may also come from a wide variety of anthropogenic sources, including combustion by cars, gas stoves, and heating appliances (National Research Council (U.S.) Committee on Toxicology, 1980). In cars, methane, ethane, and other hydrocarbons are converted into formaldehyde through the catalytic vapor phase oxidation over a metal oxide catalyst and produced by the incomplete combustion of hydrocarbons (National Library of Medicine, 1980). When methanol undergoes a chemical reaction at a high temperature, it produces formaldehyde as a byproduct (Jarvis, 2017).
Despite its many uses, exposure to formaldehyde can cause irritation to the nose, eyes, skin, and throat. Formaldehyde causes irritation in the respiratory tract with a concentration as low as 0.1 ppm when inhaled. It can cause eye irritation at concentrations of 0.05-0.10 ppm. Formaldehyde is a mucous-membrane and skin irritant and can cause conjunctivitis and lacrimation in the eyes and severe burns to the skin (Medical Management, 2014; National Research Council, 1980). Studies have shown that allergic contact dermatitis has become increasingly common since formaldehyde resins are widely used in the textile industry as an anti-wrinkle and crease-resistant component (Valdes et al., 2020).
Furthermore, prolonged exposure to formaldehyde may result in increased cancer risks and neurological dementia (Kou et al., 2022). Formaldehyde demonstrates genotoxicity, the ability to damage genetic information in cells, as it can cause increased micronucleus formation, sister chromatid exchanges, and chromosome aberrations (Kang et al., 2021). Employees in occupations that have high exposure to formaldehyde have significantly higher cancer risks: up to several thousand times higher than the limit recommended by the Environmental Protection Agency (Adamović et al., 2021). Additionally, the risk of squamous-cell carcinomas of the nasal cavities in animals and nasopharyngeal cancers in humans increases with formaldehyde exposure (Adamović et al., 2021) Formaldehyde is also a neurotoxin that affects motor functions, memory, and learning. Although formaldehyde occurs naturally, it is also formed in the process of normal metabolism in the human brain. However, exposure to high levels of formaldehyde can still severely affect metabolism, causing neurodegeneration and cognitive impairment (Tulpule & Dringen, 2013).
With its high usage and high toxicity, accurate monitoring of formaldehyde levels becomes a priority for places that work with this compound. Current methods of formaldehyde detection include mechanical detection, lab sampling, and simply smelling the chemical. However, these detection methods can be inaccurate, expensive, and hazardous, rendering them less than ideal. Most mechanical formaldehyde detectors generate false positive results when placed next to oranges (Talhelm, 2018). Laboratory testing is time-consuming and costly and smelling is direct yet unreliable. Some people only start to sense the gas at 0.117 ppm, which is above the recommended concentration. Others, such as those who smoke, are more prone to experience formaldehyde’s health effects as they have significantly higher thresholds (Berglund, 1992). After sensing increased levels of formaldehyde in an enclosed area, effective ways of remediating formaldehyde include decreasing the room temperature and humidity, which decreases rates of formaldehyde emission (Parthasarathy et al., 2011). Improving room ventilation and allowing products to off-gas before moving in are also effective ways of avoiding formaldehyde build up. (Minnesota Department of Health, 2022).
To address potential shortcomings of current detectors, we aim to construct an accurate, efficient, and cost-effective formaldehyde detector. Our study aims to utilize the detective feature of Pfrm promoters. Using a Pfrm promoter, an “ideal candidate for engineering” (Rohlhill et al.,2017), we will build a sensor that shows yellow when no formaldehyde is detected and turns red as formaldehyde builds up. The users can detect formaldehyde’s presence on furniture or other surfaces by collecting a quick swab and inserting it in the detection solution.
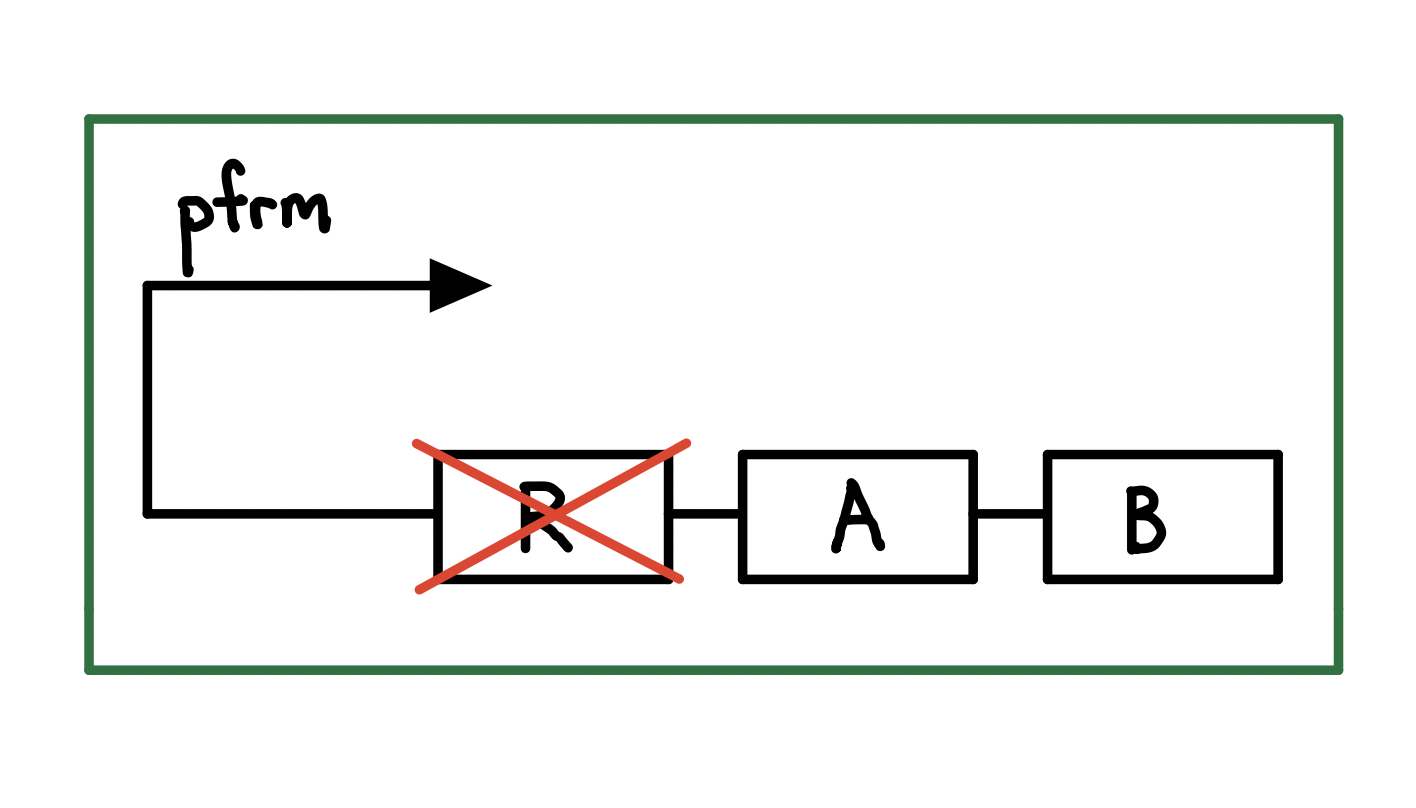
We used Pfrm because it can detect formaldehyde in the cell, and it is found in the naturally appearing formaldehyde-remediating frmRAB operon in E. coli. FrmR binds to the Pfrm promoter and inhibits its activity (Figure 1). The presence of formaldehyde negatively allosterically modulates frmR, leading to the expression enzymes FrmA and FrmB, which detoxify formaldehyde (Denby et al., 2016). The E. coli strain used in the construct is the ΔfrmR strain, which is engineered to possess a higher expression level for the genes downstream of Pfrm (Rohlhill et al., 2017).
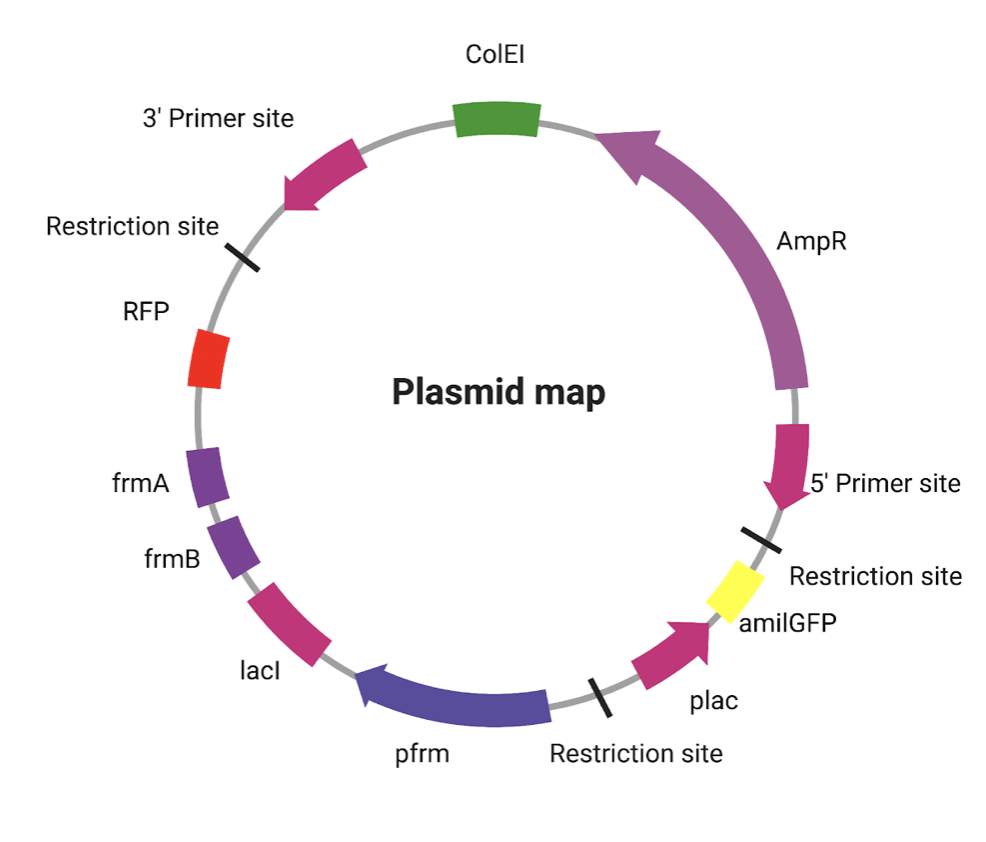
The absence of formaldehyde will trigger the yellow pigment amilGFP. Conversely, when formaldehyde is present, the promoter will activate RFP, LacI, frmA, and frmB (Figure 2). To signal the change of activation in E. coli, our group will employ a repressor system. The repressor system utilized in the construct is the Plac-lacI repressor system. PLac is a repressible constitutive promoter if cultured in lactose-rich media. Therefore, our design will grow the E. coli strain under a large supply of lactose, to ensure the activation of PLac promoter. When formaldehyde is present in the surroundings, lacI will be expressed and block the lactose promoter, stopping the expression of the amilGFP. The E. coli will further demonstrate red color instead of yellow color.
Systems level
As a solution, our detector will react to formaldehyde contents inside a swabbing device. After the user collects a sample by swabbing a target surface that they wish to test and inserting it into the detector solution, where the Pfrm promoter will be used to identify the presence of formaldehyde. If formaldehyde is absent or at levels below 100 μM, the plasmid will express a yellow pigment amilGFP (Rohlhill et al., 2017). If formaldehyde is present, the Plac-lacI repressor system will repress the originally presented yellow pigment and activate the red pigment RFP to signify the presence of formaldehyde. It will also activate frmA and frmB, which function to detoxify formaldehyde. The strain used will be the ΔfrmR. By removing FrmR, a repressor of Pfrm, from the frmRAB operon, the genes downstream of Pfrm are able to express at their maximum level (Rohlhill et al., 2017).
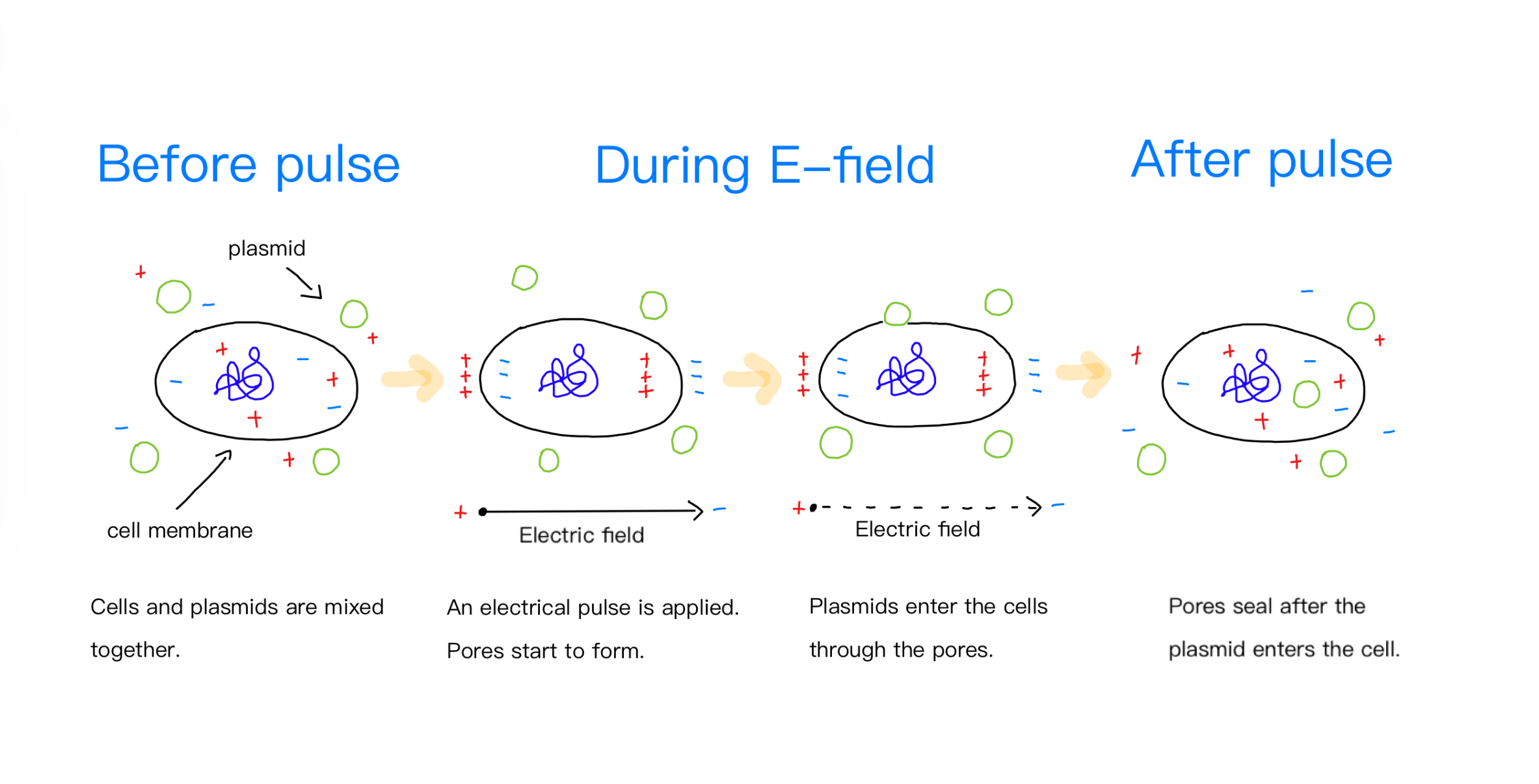
We will insert our construct onto the R0010 (pLacI)_AB plasmid backbone. The key components of the plasmid include a lac promoter, a lac operator, a biobrick prefix, a biobrick suffix, and an ampicillin resistance gene. Using restriction enzymes SpeI and PstI, we will insert the amilGFP gene (BBa_K592010) to the biobrick suffix. Using restriction enzymes EcoRI and XbaI, we will insert Pfrm, LacI, FrmA, FrmB, and RFP (BBa_E1010) to the biobrick prefix. The plasmid will be transformed into E. coli through electroporation (Figure 3).
Device level
The plasmid design contains the frmA gene, frmB gene, LacI gene, the ampicillin resistance gene, the red fluorescent protein (RFP), the naturally yellow appearing green fluorescent protein (amilGFP) from the coral Acropora millepora, the LacI repressible constitutive promoter PLac, and the formaldehyde sensing promoter Pfrm. Pfrm promotes the genes frmA, frmB, LacI, and RFP, while PLac promotes the amilGFP gene. The expression of amilGFP signifies the livelihood of E. coli prior to the exposure of formaldehyde. The RFP acts as a marker protein for the gene frmA, frmB, and LacI, which enables visual recognition when the three genes are expressed in the presence of formaldehyde.
Parts level
Our group utilizes a strain of E. coli, ΔfrmR strain edited by the lab that cuts out the frmR gene, in order to maximize the expression of the design system (Rohlhill et al., 2017). Pfrm is central to our detector. Found in the frmRAB operon in E. coli, Pfrm is a promoter that responds to formaldehyde presence. FrmR binds to the Pfrm promoter and inhibits its activity. The presence of formaldehyde negatively allosterically modulates frmR, leading to the expression of FrmA and FrmB (Denby et al., 2016). FrmA and FrmB are the enzyme factors responsible for the detoxification of formaldehyde. The enzymes encode a formaldehyde dehydrogenase, catalyzing the oxidation of formaldehyde to a less toxic compound, methanoate, which is the conjugate base of formic acid (Rohlhill et al., 2017).
Our model utilizes the Plac-LacI Repressor system to achieve the change of color in response to the presence of formaldehyde. It consists of two parts: the lactose promoter (PLac) and the lactose repressor (LacI). The lactose promoter, also
known as the lac promoter, is found in E. coli and is responsible for controlling the expression of genes involved in lactose metabolism. When a large quantity of lactose is present, PLac will activate and induce the synthesis of lactase, further expressing a color change through the AmilGFP pigment. The other vital part of this system is the lac repressor (LacI) which blocks transcription of the downstream genes without lactose. When lactose is present, it binds to LacI and causes a conformational change that allows transcription to proceed, producing enzymes needed for lactose metabolism. In our model, the lactose repressor expressed through Pfrm will act as a selection marker for the electroporation transformation.
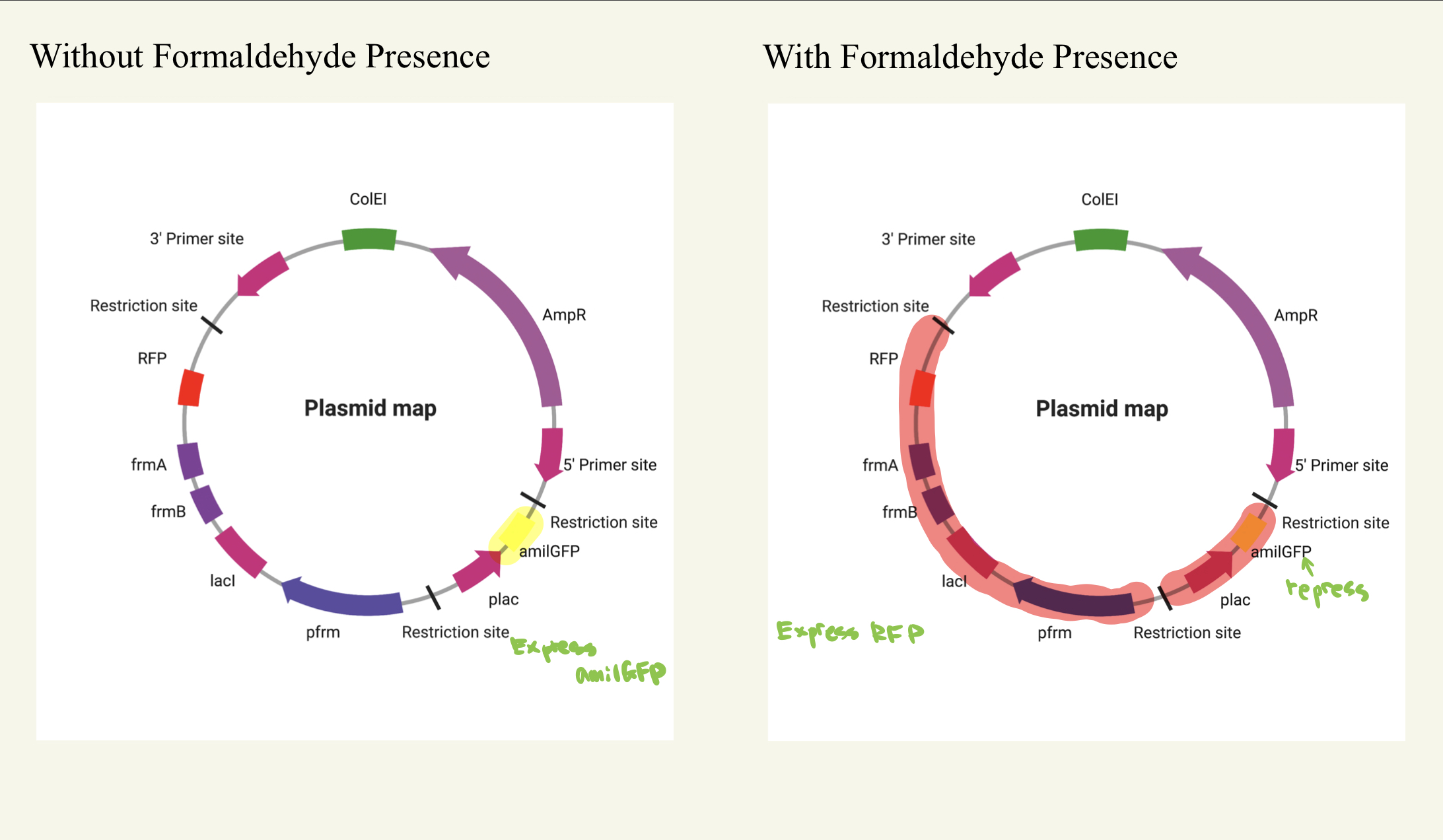
The pigment we will use in our model is AmilGFP, a yellow fluorescent protein. Derived from a coral species known as Anemonia millepora, AmilGFP naturally fluoresces bright green under blue light, allowing the track and visualization of cellular structures and processes. We will use this pigment for its ability to indicate clear and observable color changes to track the status of the detector. When the detector is armed and ready to use, AmilGFP will be expressed, indicating that there are sufficient cell activities should formaldehyde be detected and change color (Figure 4). The detector displays a red pigment to indicate the exposure to formaldehyde. We will use the red fluorescent protein (RFP) pigment, which emits red light. This pigment will be used in the system to demonstrate the presence of formaldehyde.
To initiate the gene expression downstream of Pfrm and PLac, we will employ BBa_J61100 from the Anderson RBS collection as the ribosome binding site. In addition, we will use the bidirectional double terminator BBa_B0014 to ensure the expression of only the desired genes.
We will be inserting those parts into pBR322, a plasmid commonly used in molecular biology research. We choose to use this specific plasmid since it has two biobricks that can provide an efficient approach for us to edit. In addition, the plasmid is safe, relatively simple, and has a high copy number. The plasmid contains an ampicillin resistance gene, AmpR, a protein encoded by the ampicillin resistance gene, inactivates the β-lactam ring of antibiotics and thus creates resistance against them. AmpR will act as a selection marker for the electroporation transformation.
Safety
Formaldehyde is a noxious gas with significant health effects from prolonged exposure at high concentrations. There are legal exposure limits to formaldehyde depending on the duration of exposure. The short-term exposure limit is 2 ppm for at most 15 minutes. If exposed to formaldehyde for any longer, you must not exceed an average of 0.75 ppm with a maximum exposure time of 8 hours (California Department of Health, 2011). We can measure the amount of formaldehyde in the air using a rapid test to detect the levels. When working with formaldehyde, proper PPE is the recommended safety measure to prevent intake of formaldehyde (University of California, Berkeley, 2012).
We use ΔFrmR E. coli at a Biosafety level 1 since it is non-pathogenic. For that reason, this strain does not pose any safety concerns to the human body. Our detector is generally designed to be safe around potential consumers. Because of the harmless nature of our E. coli strain, only limited precautions are necessary. In the event of detector fluid spillage, the consumer should take disinfection procedures, using isopropyl alcohol or high concentration ethanol to remove potential contamination.
Discussion
The detector can be applied to a variety of settings, such as home screening, school safety, and in manufacturing facilities. The ability to quickly collect samples through swabbing and test in detector solution makes the process of formaldehyde related disease prevention simple and effective. The reusable nature of our detector makes the process significantly less expensive compared to conventional methods of collecting sample by sample. The method of direct swabbing also allows the detector to be more accurate, where the user can pinpoint the source of formaldehyde of large concentrations.
To ensure the Plac-lacI system functions effectively and has the constitutive characteristics for the lactose promoter, we will cultivate the E. coli in a lactose-rich environment. However, a potential issue arises at low to medium concentrations of formaldehyde where lacI can be bound to the lactose present in the solution. When insufficient amounts of lacI reaches the lactose promoter, the detector will still show yellow colors despite the detection of formaldehyde. Therefore, a test will be conducted to determine the amount of lactose to be added to the solution.
Next steps
Further research should focus on finding the most effective method of formaldehyde detection. Specifically, studies should aim to clarify the levels of formaldehyde that can be quantified through the detector and the sensitivity of the detecting system. For example, using a swab to collect formaldehyde on furniture surfaces may yield inaccurate results due to the small surface area of samples taken, thus failing to detect the actual and real-time formaldehyde level in a room. Furthermore, our group will experiment on the electroporation process of the cell. However, if our construct proves capable of detecting, signaling, and remediating formaldehyde, it may have the potential to be standardized to detect other poisonous gasses such as carbon monoxide, nitrogen oxide, and sulfur dioxide. The key is to identify the corresponding promoter. For example, a simple switch from Pfrm to CooA, a CO-sensing transcriptional activator (Techtmann et al., 2011), may turn the construct into a carbon monoxide detector.
Author contributions
L.D. is the founder of the project. She researched and wrote on formaldehyde influences, and is also responsible for the art design, creation, and voice over the video. B.J researched the pigment genes and the promoters and is responsible for the construct design. He wrote the script for the promotional video. J.M studied the safeties for the project using formaldehyde and also studied the origins of formaldehyde in the common world. E.T is the main artist for the project. He designed the plasmid and assisted in the explanation for the background of the project. G.W. contributes to the art section of the project and is also responsible for the research on the system. M.Y is the main editor and script writer of the paper, he also researched the current methods and explored advantages the project has over various industries.
Acknowledgements
We would like to thank Dr. Beth Pethel for her continuous support and guidance throughout this project.
References
Adamović, D., Čepić, Z., Adamović, S., Stošić, M., Obrovski, B., & Morača, S. (2021). Occupational exposure to formaldehyde and cancer risk assessment in an anatomy laboratory. International Journal of Environmental Research and Public Health. https://doi.org/10.3390/ijerph182111198
Agency for Toxic Substances and Disease Registry. (2014, October 21). Medical management guidelines for formaldehyde. https://wwwn.cdc.gov/TSP/MMG/MMGDetails.aspx?mmgid=216&toxid=39
Berglund, B., & Nordin, S. (1992). Detectability and perceived intensity for formaldehyde in smokers and non-smokers. Chemical Senses, 17(3), 291-306. https://doi.org/10.1093/chemse/17.3.291
Bernstein, R. S., Stayner, L. T., Elliott, L. J., Kimbrough, R., Falk, H., & Blade, L. (1984). Inhalation exposure to formaldehyde: An overview of its toxicology, epidemiology, monitoring, and control. American Industrial Hygiene Association Journal, 45(11), 778-785. https://doi.org/10.1080/15298668491400601
Centers for Disease Control and Prevention. (2016, September 18). Chemical disinfectants. https://www.cdc.gov/infectioncontrol/guidelines/disinfection/disinfection-methods/chemical.html
Denby, K. J., Iwig, J., Bisson, C., Westwood, J., Rolfe, M. D., Sedelnikova, S. E., Higgins, K., Maroney, M. J., Baker, P. J., Chivers, P. T., & Green, J. (2016). The mechanism of a formaldehyde-sensing transcriptional regulator. Scientific Reports, 6(1), 38879. https://doi.org/10.1038/srep38879
EH&S. (2012, August 1). Formaldehyde: Hazards and precautions. https://ehs.berkeley.edu/sites/default/files/publications/formaldehyde-fact-sheet.pdf
Ferenc, M. (2017, November). Plasmids 101: Common lab E. coli strains. Addgene blog. https://blog.addgene.org/plasmids-101-common-lab-e-coli-strains
Herrera-rodriguez, J., Signorazzi, A., Holtrop, M., De vries-idema, J., & Huckriede, A. (2019). Inactivated or damaged? Comparing the effect of inactivation methods on influenza virions to optimize vaccine production. Vaccine, 37(12), 1630-1637. https://doi.org/10.1016/j.vaccine.2019.01.086
Jarvis, D. L. (2017). Polyacetals. Brydson’s Plastics Materials, 513-526. https://doi.org/10.1016/B978-0-323-35824-8.00019-0
Kang, D. S., Kim, H. S., Jung, J. H., Lee, C. M., Ahn, Y. S., & Seo, Y. R. (2021). Formaldehyde exposure and leukemia risk: A comprehensive review and network-based toxicogenomic approach. Genes and Environment, 43(1). https://doi.org/10.1186/s41021-021-00183-5
Kou, Y., Zhao, H., Cui, D., Han, H., & Tong, Z. (2022). Formaldehyde toxicity in age-related neurological dementia. Ageing Research Reviews, 73, 101512. https://doi.org/10.1016/j.arr.2021.101512
Mccrate, O. A., Zhou, X., & Cegelski, L. (2013). Curcumin as an amyloid-indicator dye in E. coli. Chemical Communications, 49(39), 4193. http://doi.org/10.1039/c2cc37792f
Minnesota Department of Health. (2022, October 3). Formaldehyde in your home. https://www.health.state.mn.us/communities/environment/air/toxins/formaldehyde.htm
National Institute of Environmental Health Sciences. (2023, January 3). Formaldehyde. Retrieved January 17, 2023, from https://www.niehs.nih.gov/health/topics/agents/formaldehyde/index.cfm
National Research Council. (1980). Formaldehyde-An assessment of its health effects. National Academies Press. https://doi.org/10.17226/705
Occupational Safety and Health Administration. (n.d.). 1910.1048 – Formaldehyde. https://www.osha.gov/laws-regs/regulations/standardnumber/1910/1910.1048
Osman, D., Piergentili, C., Chen, J., Sayer, L. N., Usón, I., Huggins, T. G., Robinson, N. J., & Pohl, E. (2016). The effectors and sensory sites of formaldehyde-responsive regulator frmr and metal-sensing variant. Journal of Biological Chemistry, 291(37), 19502-19516. https://doi.org/10.1074/jbc.M116.745174
Parthasarathy, S., Maddalena, R. L., Russell, M. L., & Apte, M. G. (2011). Effect of temperature and humidity on formaldehyde emissions in temporary housing units. Journal of the Air & Waste Management Association, 61(6), 689-695. https://doi.org/10.3155/1047-3289.61.6.689
Rohlhill, J., Sandoval, N. R., & Papoutsakis, E. T. (2017). Sort-seq approach to engineering a formaldehyde-inducible promoter for dynamically regulated Escherichia coli growth on methanol. ACS Synthetic Biology, 6(8), 1584-1595. https://doi.org/10.1021/acssynbio.7b00114
Shimomura, T. (2014). Nanoporous silicon biochemical sensors. Nanosensors for Chemical and Biological Applications, 254-266. https://doi.org/10.1533/9780857096722.2.254
Techtmann, S. M., Colman, A. S., Murphy, M. B., Schackwitz, W. S., Goodwin, L. A., & Robb, F. T. (2011). Regulation of multiple carbon monoxide consumption pathways in anaerobic bacteria. Frontiers in Microbiology, 2. https://doi.org/10.3389/fmicb.2011.00147
Tulpule, K., & Dringen, R. (2013). Formaldehyde in brain: An overlooked player in neurodegeneration? Journal of Neurochemistry, 127(1), 7-21. https://doi.org/10.1111/jnc.12356
United States Environmental Protection Agency. (2022, April 18). Facts about formaldehyde. Retrieved December 15, 2022, from https://www.epa.gov/formaldehyde/facts-about-formaldehyde
United States Environmental Protection Agency. (2022, April 22). Frequent questions for consumers about the formaldehyde standards for composite wood products act. https://www.epa.gov/formaldehyde/frequent-questions-consumers-about-formaldehyde-standards-composite-wood-products-act
U.S. Food and Drug Administration. (2019, April 19). Common Ingredients in U.S. Licensed Vaccines. https://www.fda.gov/vaccines-blood-biologics/safety-availability-biologics/common-ingredients-us-licensed-vaccines
Valdes, F., Mcnamara, S. A., & Keri, J. (2020). Allergic contact dermatitis from transient formaldehyde exposure in a traveler: Are all backpacks created equal? Cureus,12(12). https://doi.org/10.7759/cureus.12252