Logan Amos, Lujiong Danzhen, Phalak Dhingra, Erika Frohring, Isabella Haslinger Johnson, Sofia Kovacevic, Danyan Li, Chongming Liu, Yihan Ma, Anya Mathur, Sheilla Muligande, Annie Nguyen, Ruihan Qin, Carmen Reed, Aparajita Shimpi, Lingzhen Wang, Cavin Xue, Siwen Yang, Mira Zamarro, and Yuejiao (Vivian) Zhou ▫ Western Reserve Academy, Hudson OH
Reviewed on 6 May 2023; Accepted on 13 July 2023; Published on 16 October 2023
With help from the 2023 BioTreks Production Team.
Dental caries is a primary oral health concern, affecting 90% of adults worldwide. Invasive treatments such as dental fillings and root canal therapies stop the degrading effects of caries and accelerate the remineralization process. Although current solutions are effective, they can be painful and expensive for patients. Additionally, these treatments are temporary and may require repetitive maintenance. Streptococcus mutans, a naturally occurring bacterial strain in the human oral cavity, is a principal pathogenic agent of caries and tooth decay. Saliva acidity and accumulation of food debris create an ideal environment for S. mutans proliferation. The bacterium becomes part of a biofilm on the surface of teeth, catalyzing plaque production. Studies demonstrate inhibition of the virulence and proliferation of S. mutans by Streptococcus salivarius A12. Researchers are exploring the possibility of using these bacteria to combat dental caries. Inspired by these findings, our project aims to forestall dental caries by using S. salivarius M18, another bacterial strain in the oral cavity that produces two enzymes, urease and dextranase. These enzymes break down dental plaque and decrease saliva acidity. M18 effectively minimizes tooth enamel erosion and impedes dental caries formation by disrupting biofilm formation and reducing salivary pH. We will transform the genes ureA-C and dex, encoding enzymes urease and dextranase, into an early enamel colonizer, Rothia aeria, to suppress caries. Our product will be a R. aeria paste applied after dental prophylaxis to forestall dental caries formation. This research demonstrates a biosynthetic method that improves current dental treatments and designs a biocompatible product for oral hygiene.
Keywords: Streptococcus mutans, dental caries, Streptococcus salivarius M18, urease, dextranase, caries prevention, probiotics
Authors are listed in alphabetical order. Beth Pethel from Western Reserve Academy, Hudson OH mentored the group. Please direct all correspondence to pethelb@wra.net.
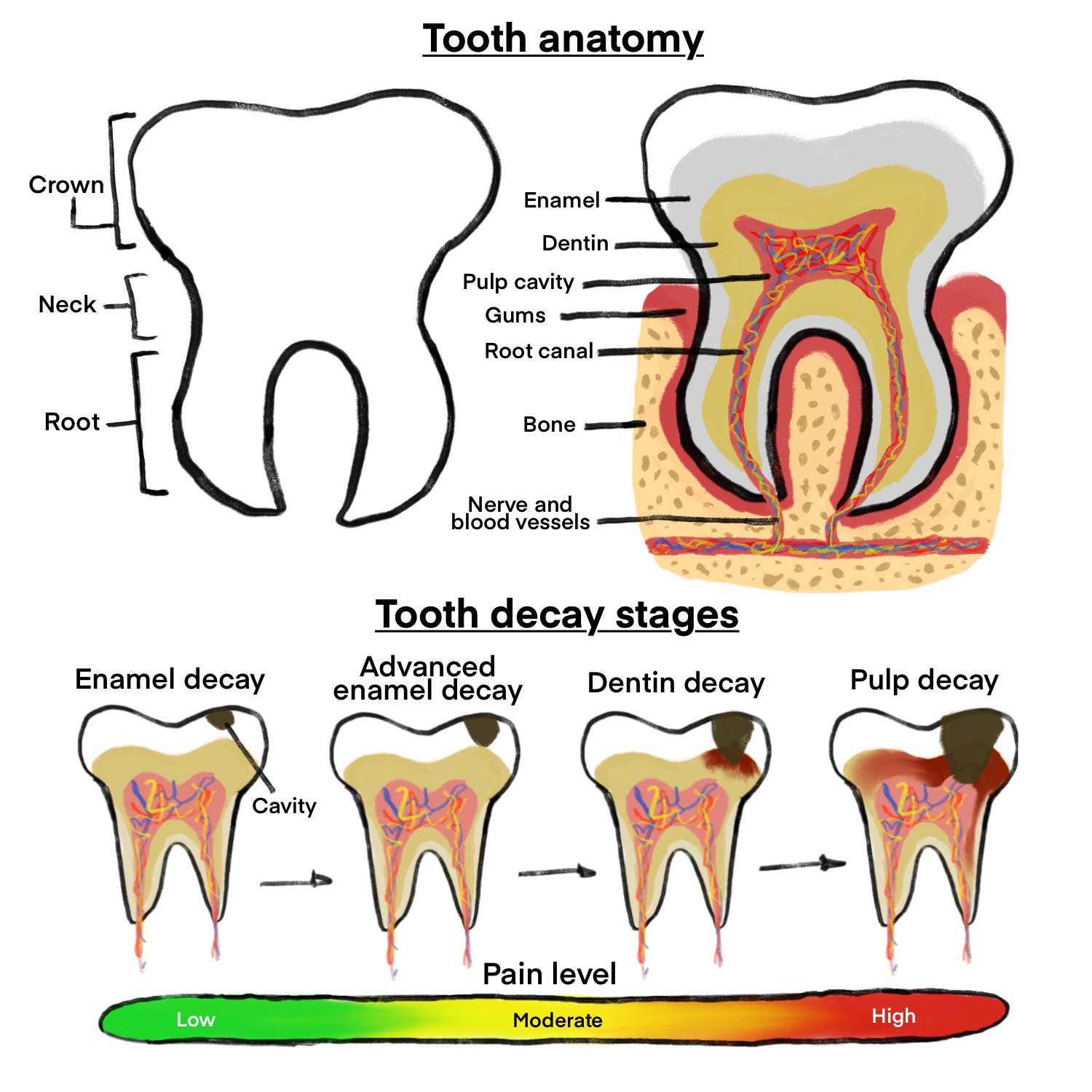
Dental caries, also known as cavities or tooth decay, affects approximately 90% of adults ages 20-64. The primary cause of tooth decay is plaque, a sticky film of bacteria that forms on the tooth’s surface and remains present for extended periods, as seen in Figure 1. Several factors can contribute to plaque production but the most notable element is the overabundance of sugars in processed foods and drinks (Dental Plaque, n.d.). Plaque, in addition to microorganisms and food debris, contributes to the formation of dental cavities (Forssten et al., 2010, p. 291). Plaque-colonizing bacteria, such as the acid-producing S. mutans, damage the tooth structure, especially near carbohydrates such as sucrose or fructose (Forssten et al., 2010, p. 291). Tooth decay may result in tooth loss, tissue damage, or potentially life-threatening issues (Cavities/tooth Decay, 2022).
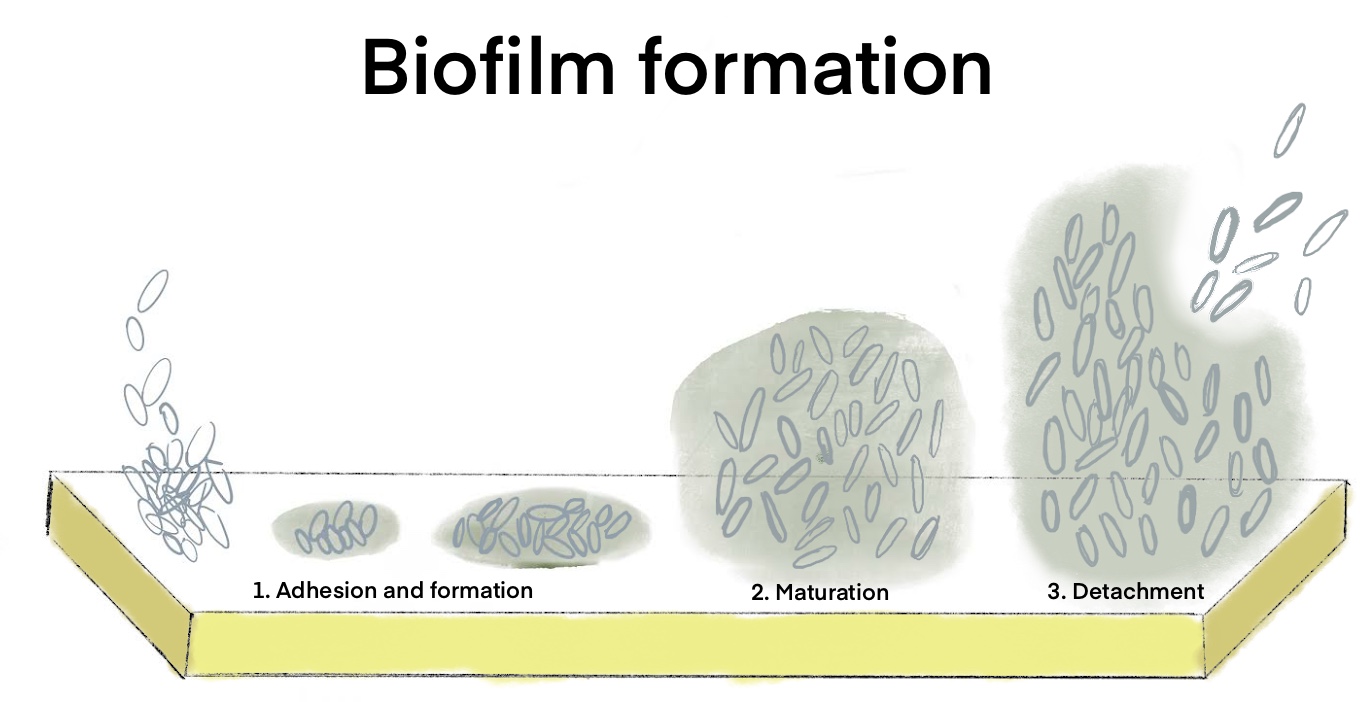
Biofilms, layers of bacteria that adhere to a surface (Figure 2), are abundant throughout the mouth. Biofilms protect teeth by maintaining a balance between harmless and destructive bacteria (Brown, 2022). However, they also contribute to plaque production and subsequent tooth decay, as seen in Figure 1. Although proper dental hygiene prevents the continuous growth of biofilms and reduces the risk of cavities, the biofilm still reforms four hours after its removal (Dhir, 2013).
As plaque reforms on clean teeth, certain organisms, known as early colonizers, repopulate the mouth before others. S. mutans, one of these early colonizers, is one of the main contributors to tooth decay and plaque formation (Lemos et al., 2019). After binding to the tooth and cultivating a biofilm, S. mutans synthesizes dextran (MicrobeWiki, 2016), a common sugar produced by bacteria (Redasani & Bari, 2015).
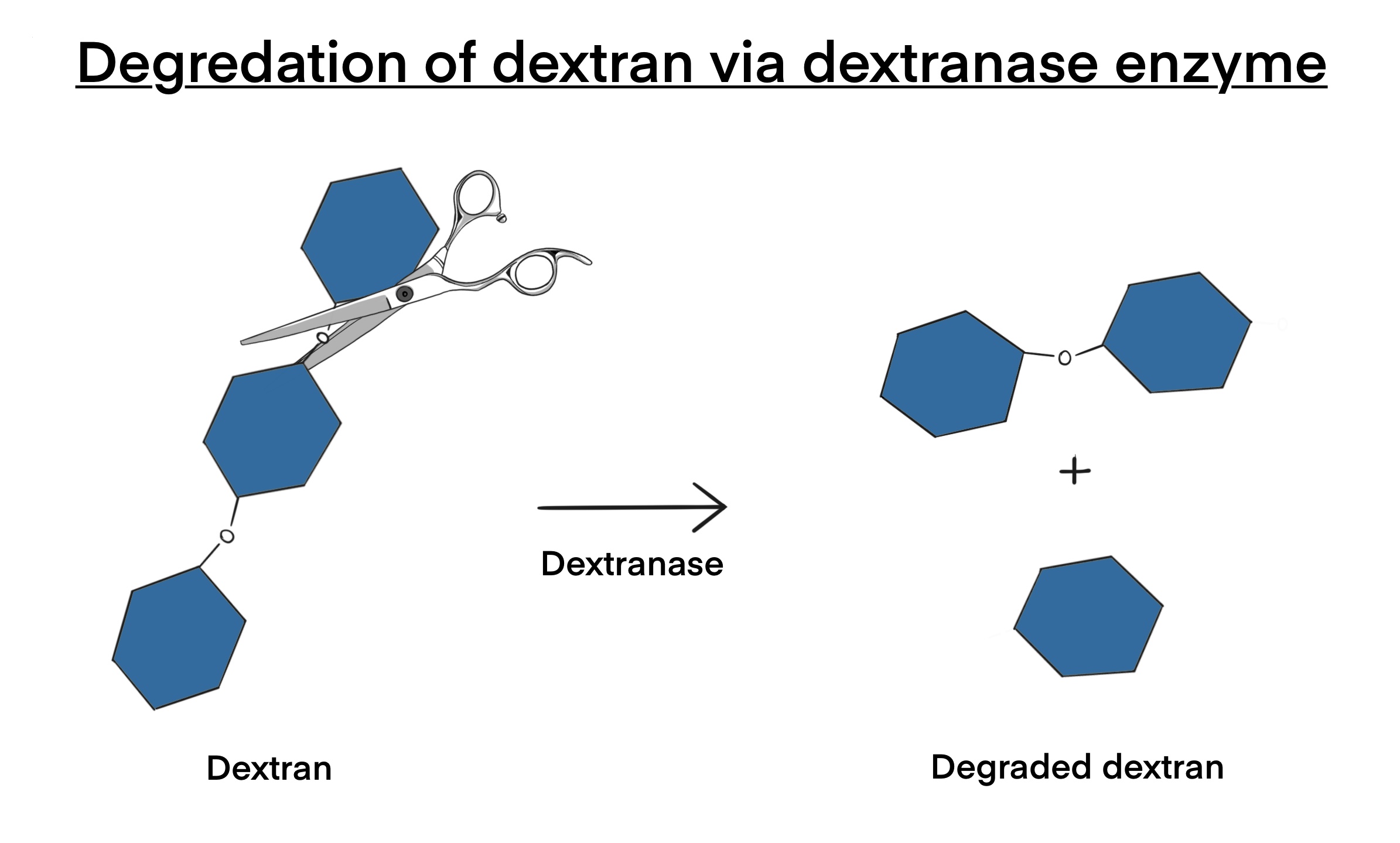
Another early colonizer of the mouth’s soft tissue, S. salivarius M18, produces the enzymes dextranase and urease. Dextranase breaks down dextran in the plaque of decaying teeth (Burton et al., 2013). Bacteria such as S. mutans synthesize dextran (MicrobeWiki, 2016). Dextran is found in beets and sugarcane (Infinita Biotech, 2022). Dextranase regulates the mildly toxic polysaccharide dextran levels in the mouth (Infinita Biotech, 2022). Through its breakdown of sugar, dextranase helps neutralize plaques that contribute to tooth decay (Stowik, 2016, p. 16).
The enzyme urease uses the compound urea to generate ammonia, which neutralizes certain acids in the mouth that may cause tooth decay (Chen et al., 2000, p. 4667). Extended exposure to acidic environments contributes to increased growth of harmful bacteria, and a pH value below 5.5 in the oral environment is a critical factor in the progression of cavity development (Matsui & Cvitkovitch, 2010). Urea is a naturally occurring compound commonly found in saliva and urine. It is also used in agriculture due to its tendency to form pH-heightening ammonia and decrease acidity (American chemical society, 2021). Urease hydrolyzes the compound, utilizing its high nitrogen content to generate the alkali ammonia (Chen et al., 1996, p. 585). In doing so, it neutralizes acids created by dietary carbohydrates (Morou-bermudez et al., 2011, p. 1283). Urease is made up of three subunits that are encoded by the genes ureA-C. It is also one of the few enzymes that require a nickel ion to function. Consequently, the inclusion of the genes ureDEFG is present for the production of functional urease.
This project aims to insert the dextranase seen in Figure 3 and urease genes from S. salivarius into the early colonizer R. aeria. R. aeria is an effective carrier for the genes, as it colonizes the biofilms of teeth while S. salivarius only inhabits the oral tissue. R. aeria, similar to S. salivarius, possesses a Biosafety Level 1 rating and is gram-positive (Rothia aeria, n.d.).
Systems level
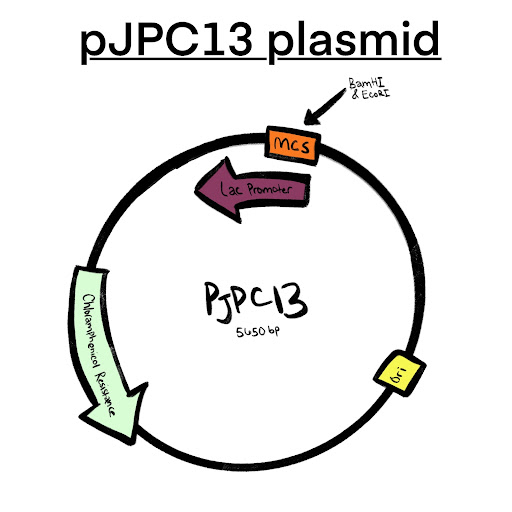
The goal of this research is to develop a paste that contains enzymes for use in the oral cavity. To achieve this, we are introducing four genes (ureA, ureB, ureC, and dex) that encode for urease and dextranase enzymes into R. aeria. To do this, we will insert each of the target genes into a plasmid called pJPC13, seen in Figure 4, which includes a lac promoter, a lac terminator, and a chloramphenicol resistance marker. We will then transform the plasmid into R. aeria, which will be grown with Medium 92 containing chloramphenicol to isolate a single colony for inoculation. By growing the bacteria on chloramphenicol agar plates, we will ensure that only successfully transformed bacteria form colonies. As we do not need to extract the transformed bacteria, the final product will be the active R. aeria colonies in a paste. While the plasmid we are using is not specifically designed for R. aeria, our team has chosen a plasmid that is designed for a bacterium that genetically mimics it based on limited research.
Device level
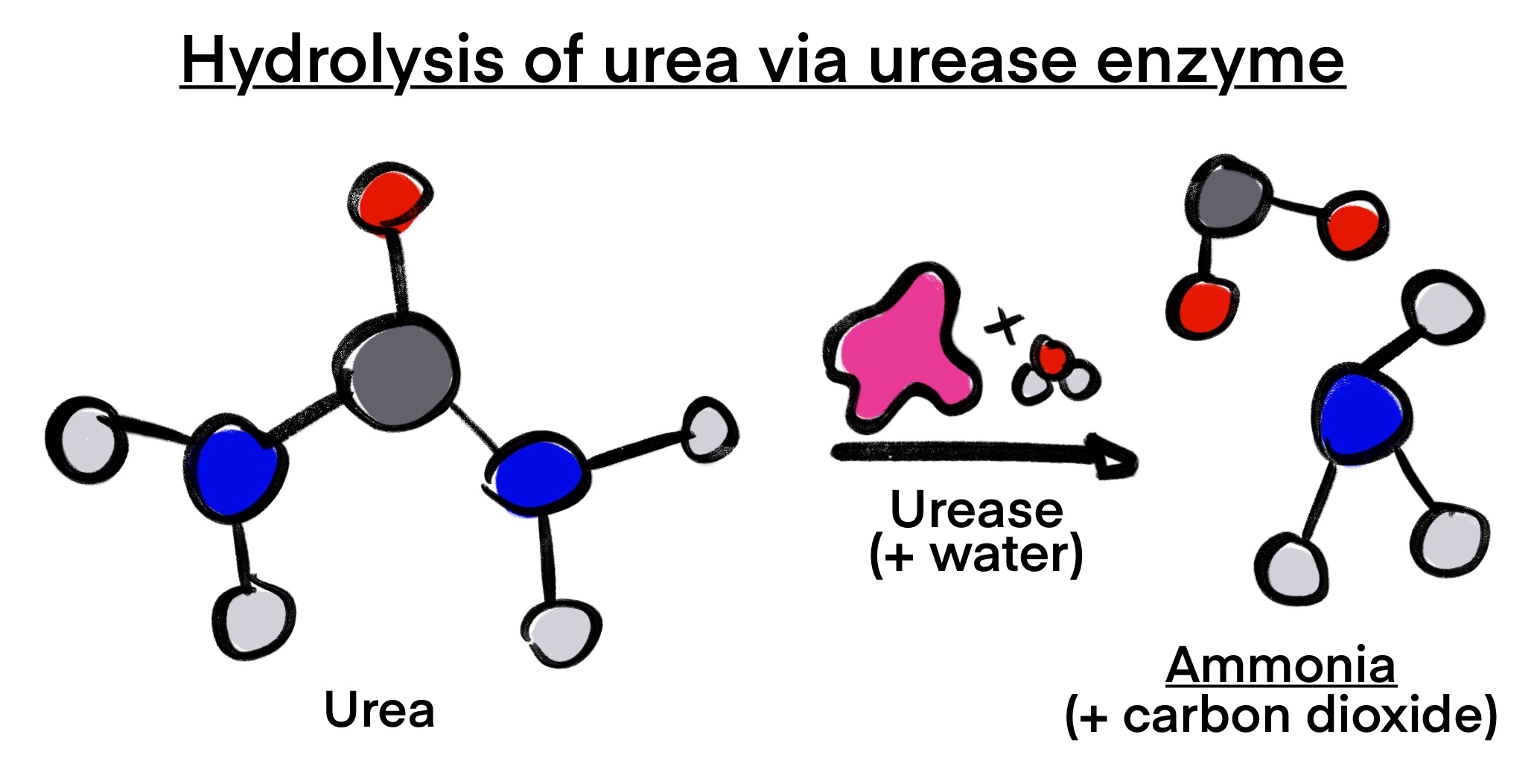
Our project utilizes Rothia aeria A 1-17B as the chassis, primarily because it is an early colonizer of human teeth and is expected to have a minimal impact on the oral microbiome. We will use the ureA-C gene cluster, which encodes for the enzyme urease, and the dex gene, which encodes for dextrase. Dextrase is an enzyme that breaks down dextrane sugar, converting it into a non-toxin. Specifically, we only require ureA, B, and C to produce urease. Urease catalyzes the reaction between urea and water, producing ammonia and carbon dioxide, as seen in Figure 5. Ammonia then increases the pH of the oral cavity, thus creating an environment that is not optimal for the growth of S. mutans. In addition, we will use the plasmid backbone pJPC13, which contains a lac promoter, a lac terminator, and a chloramphenicol resistance marker.
Parts level
Our project focuses on creating a paste containing dextranase and urease enzymes commonly found in S. salivarius. S. salivarius is a probiotic bacterium that can reduce oral infections by producing bacteriocin-like inhibitory substances. We plan to use the dex gene to produce dextranase and the ureA-C genes to produce urease. To simplify the process, we will use the lac promoter and lac terminator, which are present in the pJPC13 plasmid and are also related to lactose that is found in the oral cavity. The BamH1 and EcoR1 restriction enzymes present in the multiple cloning site of the pJPC13 plasmid will be used to insert the selected genes. As these enzymes occur only once in the multiple cloning site, we can avoid cleavage during insertion. Our selectable marker will be chloramphenicol, as R. aeria is not naturally resistant to this antibiotic.
Safety
The safety of R. aeria must be considered in topical use. The A 1-17B strain of R. aeria is classified as a Biosafety Level 1 organism and presents minimal risk to healthy humans (Strain Identifier, 2022; Wei et al., 2020, p 1). In some cases, R. aeria has been found at sites of infective endocarditis; however, the oral presence and consumption of the bacteria were not considered a direct cause (Nicodemo et al., 2014). The growth medium used for R. aeria cells, Medium 92, is categorized as Biosafety Level 1 (DSMZ, n.d.), meaning the media poses no hazards and does not require further safety measures. The electroporation technique this project will use to transform R. aeria poses no safety concerns.
In this project’s R. aeria treatment, urease production remains a concern. Urease has been shown to have toxic effects on human cells and can cause possible skin and eye irritation due to its enzymatic activity (Konieczna et al., 2012, p. 789). However, ammonium ions in the growth medium can limit urease transfer across the cell membrane, and glucose-inhibited enzyme synthesis can further regulate its production (Magaña-plaza & Ruiz-herrera, 1967, p. 1294). Dextranase is safe for human consumption, and allergic reactions are rare (Silano et al., 2020, p. 6309).
Discussion
This project aims to prevent dental caries using S. salivarius M18, another bacterial strain in the oral cavity that produces two enzymes, urease, and dextranase. By disrupting the natural formation of biofilm and reducing salivary pH, M18 should protect the teeth against enamel erosion and the formation of dental caries. By transforming the ureA-C and dex genes into Rothia aeria, a paste of the transformed R. aeria could be applied during regular dental visits to prevent cavities.
However, this method may present potential hazards during practical use. Consultation with professionals and safety experiments would confirm that additional levels of R. aeria in the oral cavity do not cause adverse effects. As outlined in the safety section, its use poses minimal risk. The electroporation technique and use of dextranase are both safe for use in this project. While urease may cause adverse effects, the process of glucose-inhibited enzyme synthesis should neutralize these effects. If this project proves successful, the product could be used as a widespread preventative against plaque and dental caries.
Next steps
Though this project remains theoretical, testing must be considered in the laboratory environment. We will begin with growing Rothia aeria. To prepare for DNA extraction, we will make a stock solution of chloramphenicol and use it to isolate bacterial colonies. Since the bacterium is aerobic, an oxygen-free environment is not required for growth (Taira & Aoki, 2019). Growth will occur on Medium 92 at body temperature (37°C) to simulate the oral environment. Then, we will use electroporation to transform R. aeria. We will then apply our transformed bacteria to lab-grown plaque originating from teeth impacted by caries. This process will be repeated using different concentrations of the bacterial product to gauge the levels of each reaction in relation to the breakdown of plaque samples. Using a previously researched system for in vitro studies of the oral cavity, we will also test adequate quantities of the dextranase and urease enzymes on already formed biofilms that contain S. mutans (Mira et al., 2019). Through this procedure, we will compare the formation of biofilm in an environment that contains urease and dextranase to an environment that does not. Clinical trials can be conducted in small groups of dental providers and patients. If the project shows a reduction in the formation of dental caries, we can seek approval from dental associations to distribute the product to a larger group of dental patients. Ideally, this project will become a prevalent dental treatment for all.
Author contributions
L.W. came up with the original idea and began introductory research. L.A., E.F., I.H., D.L., R.Q., L.W., and S.Y. conducted the early research process. L.A., L.D., P.D., E.F, I.H., S.K., D.L., C.L., Y.M., A.M., S.M., A.N., R.Q., C.R., A.S., L.W., C.X., S.Y., M.Z., and Y.Z. contributed to the writing and proofreading of the paper. P.D., E.F., and R.Q. designed the images and graphics for this project.
Acknowledgements
We would like to acknowledge Western Reserve Academy for its support and resources. We would especially like to thank our mentor, Dr. Pethel, for introducing the team to synthetic biology and for her guidance and continuous encouragement throughout the project.
References
American chemical society. (2021, January 11). Urea. https://www.acs.org/molecule-of-the-week/archive/u/urea.html
Brown, S. (2022, April 19). Can probiotics help with gum disease? WebMD. https://www.webmd.com/oral-health/probiotics-gum-disease
Burton, J. P., Drummond, B. K., Chilcott, C. N., Tagg, J. R., Thomson, W. M., Hale, J. D. F., & Wescombe, P. A. (2013). Influence of the probiotic Streptococcus salivarius strain m18 on indices of dental health in children: A randomized double-blind, placebo-controlled trial. Journal of Medical Microbiology, 62(6), 875-884. https://doi.org/10.1099/jmm.0.056663-0
Mayo Clinic. (2022, March 19). Cavities/tooth decay https://www.mayoclinic.org/diseases-conditions/cavities/symptoms-causes/syc-20352892
Chen, Y. Y., Clancy, K. A., & Burne, R. A. (1996). Streptococcus salivarius urease: Genetic and biochemical characterization and expression in a dental plaque streptococcus. Infection and Immunity, 64(2), 585-592. https://doi.org/10.1128/iai.64.2.585-592.1996
Chen, Y. Y. M., Weaver, C. A., & Burne, R. A. (2000). Dual functions of Streptococcus salivarius urease. Journal of Bacteriology, 182(16), 4667-4669. https://doi.org/10.1128%2Fjb.182.16.4667-4669.2000
Cleveland Clinic. (n.d.). Dental plaque. https://my.clevelandclinic.org/health/diseases/10953-plaque
Dhir, S. (2013). Biofilm and dental implant: The microbial link. Journal of Indian Society of Periodontology, 17(1), 5. https://doi.org/10.4103/0972-124X.107466
DSMZ. (n.d.). Rothia aeria DSM 14556. (n.d.). https://www.dsmz.de/collection/catalogue/details/culture/DSM-14556
Forssten, S. D., Björklund, M., & Ouwehand, A. C. (2010). Streptococcus mutans, caries and simulation models. Nutrients, 2(3), 290-298. https://doi.org/10.3390/nu2030290
Infinita Biotech. (n.d.). Applications of dextranase enzyme. Infinita Biotech. https://infinitabiotech.com/blog/applications-of-dextranase-enzyme/
Konieczna, I., Zarnowiec, P., Kwinkowski, M., Kolesinska, B., Fraczyk, J., Kaminski, Z., & Kaca, W. (2012). Bacterial urease and its role in long-lasting human diseases. Current Protein and Peptide Science, 13(8), 789-806. https://doi.org/10.2174%2F138920312804871094
Lemos, J. A., Palmer, S. R., Zeng, L., Wen, Z. T., Kajfasz, J. K., Freires, I. A, Abranches, J., & Brady, L.J. (2019). The biology of Streptococcus mutans. Microbiology Spectrum, 7(1). https://doi.org/10.1128/microbiolspec.GPP3-0051-2018
Magaña-plaza, I., & Ruiz-herrera, J. (1967). Mechanisms of regulation of urease biosynthesis in Proteus rettgeri. Journal of Bacteriology, 93(4), 1294-1301. https://doi.org/10.1128%2Fjb.93.4.1294-1301.1967
Matsui, R., & Cvitkovitch, D. (2010). Acid tolerance mechanisms utilized by Streptococcus mutans. Future Microbiology, 5(3), 403-417. https://doi.org/10.2217/fmb.09.129
MicrobeWiki. (2016, February 11). Streptococcus mutans– Tooth decay. https://microbewiki.kenyon.edu/index.php/Streptococcus_mutans-_Tooth_Decay
Mira, A., Buetas, E., Rosier, B. T., Mazurel, D., Villanueva-castellote, Á., Llena, C., & Ferrer, M. D. (2019). Development of an in vitro system to study oral biofilms in real time through impedance technology: Validation and potential applications. Journal of Oral Microbiology, 11(1), 1609838. https://doi.org/10.1080%2F20002297.2019.1609838
Morou-bermudez, E., Elias-boneta, A., Billings, R.j., Burne, R.a., Garcia-rivas, V., Brignoni-nazario, V., & Suarez-perez, E. (2011). Urease activity in dental plaque and saliva of children during a three-year study period and its relationship with other caries risk factors. Archives of Oral Biology, 56(11), 1282-1289. https://doi.org/10.1016%2Fj.archoralbio.2011.04.015
Nicodemo, A. C., Gonçalves, L. G., Odongo, F. C. A., Martino, M. D. V., & Sampaio, J. L. M. (2014). Rothia aeria endocarditis in a patient with a bicuspid aortic valve: Case report. The Brazilian Journal of Infectious Diseases, 18(5), 561-564. httpsdoi.org/10.1016/j.bjid.2014.05.001
Redasani, V. K., & Bari, S. B. (2015). Approaches for Prodrugs. In V. K. Redasani & S. B. Bari (Eds.), Prodrug design: Perspectives, approaches and applications in medicinal chemistry (pp. 33-49). Academic Press. https://doi.org/10.1016/B978-0-12-803519-1.00004-0
Silano, V., Barat baviera, J. M., Bolognesi, C., Cocconcelli, P. S., Crebelli, R., Gott, D. M., Grob, K., Lambré, C., Lampi, E., Mengelers, M., Mortensen, A., Rivière, G., Steffensen, I., Tlustos, C., Van loveren, H., Vernis, L., Zorn, H., Andryszkiewicz, M., Gomes, A., . . . Chesson, A. (2020). Safety evaluation of the food enzyme dextranase from collariella gracilis strain atcc‐16153. EFSA Journal, 18(12). https://doi.org/10.2903%2Fj.efsa.2020.6309
Stowik, T. A. (2016). Contribution of probiotics Streptococcus salivarius strains K12 and M18 to oral health in humans: A review [Master’s thesis, University of Connecticut]. University of Connecticut. https://opencommons.uconn.edu/cgi/viewcontent.cgi?article=1484&context=srhonors_theses
Strain Identifier. (2022, December 7). Rothia aeria. BacDive. https://bacdive.dsmz.de/strain/7780
Taira, Y., & Aoki, Y. (2019). Tube-ovarian abscess caused by Rothia aeria. BMJ Case Reports, 12(8), e229017. http://dx.doi.org/10.1136/bcr-2018-229017 Wei, G., Darwish, G., Oppenheim, F. G., Schuppan, D., & Helmerhorst, E. J. (2020). Commensal bacterium Rothia aeria degrades and detoxifies gluten via a highly effective subtilisin enzyme. Nutrients, 12(12), 3724. https://doi.org/10.3390/nu12123724