Leslie Baker, Cate Lynch, Rosalia Reale, and Hana Shinzawa ▫ BioBuilderClub, Montrose School, Medfield, Massachusetts, United States
Reviewed on 6 May 2023; Accepted on 13 July 2023; Published on 16 October 2023
With help from the 2023 BioTreks Production Team.
Salmonella typhimurium is a bacterial species that is prevalent and causes disease in sub-Saharan Africa, South Asia, East Asia, and Pacific regions. Better detection of the microbe would improve food safety and security in these regions. Because S. typhimurium is mainly present in the food and water of developing countries, an effective detector must be portable and self-regenerating. To achieve these requirements, we designed a detection system using synthetic biology. Our system will express the a-agglutinin complex subunits (Aga1p and Aga2p) on the surface of Saccharomyces cerevisiae, a yeast that is both amenable to engineering and safe to provide to households in affected areas. The antibodies presented on these engineered yeast will bind to S. typhimurium, stimulating a proton pump in the yeast and adding H+ ions to the yeast growth media by means of a signal transduction pathway not yet identified. A pH indicator, such as Bromophenol blue, will be used to detect the increase of H+ ions in solution. If the solution drops significantly below 5.75, the typical pH of S. cerevisiae, to about 4.5, the solution will turn yellow, indicating the presence of S. typhimurium. Our approach, if successful, will create a low-cost, self-regenerating system for the detection of S. typhimurium. The solution’s low cost and capacity to reproduce will ensure greater access to our system, allowing those in developing countries to discern if their food and water is safe, which will help reduce disease.
Keywords: Salmonella typhimurium detection, synthetic biology detector, yeast-based detector
Authors are listed in alphabetical order. Alexis Dimitriou, and Sarah Hanna from BioBuilderClub, Montrose School, Medfield, Massachusetts, United States mentored the group. Please direct all correspondence to .
Salmonella typhimurium (S. typhimurium) is a bacterial species that causes typhoid fever. The disease was first recognized in the 1800s and considered a worldwide issue until the mid-twentieth century when developed countries made improvements to sanitation and hygiene. With these advancements, developed countries continue to have low incidence of typhoid fever; most cases occur when travelers visit regions where typhoid fever is endemic. Areas such as North America and Europe are seen as low-incidence areas that have less than ten cases per 100,000 per year (Colomba et al, 2008). S. typhimurium is prevalent and commonly causes disease in areas such as sub-Saharan Africa, South Asia, East Asia, and Pacific regions (Als.et al, 2018).
S. typhimurium can cause sepsis and death if not treated with the correct antibiotics. However, in developing countries, where S. typhimurium bacteria is most prevalent due to the lack of access to safe water and sanitation, access to these antibiotics is limited. Better detection of the microbe would improve food safety and security in these regions. Thus, we aim to create a solution that can detect S. typhimurium in order to prevent potentially deadly infections in vulnerable populations.
S. typhimurium is most commonly found in areas where communities do not have access to clean water and sanitation. S. typhimurium is thought to affect approximately 22 million people per year and causes up to 200,000 deaths, mostly in children (Pollard, 2022). The regions of South Asia, sub-Saharan Africa, East Asia, and the Pacific generally have the highest incident rate, and S. typhimurium is considered endemic in these regions (Als et al, 2018). These areas also often lack the antibiotics to treat S. typhimurium, leading to higher mortality rates (Barrett & Stanberry, 2009). Additionally, the emerging resistance to first line antimicrobials increases treatment costs and deaths from the infection (Saporito, 2017).
Some bacteria, including S. typhimurium spreads most commonly through the feces of infected individuals. These individuals with S. typhimurium can become asymptomatic carriers, who remain contagious for years (Vyas, 2021). Thus, proper sanitation is especially important in controlling the spread of S. typhimurium. Water chlorination and sand filtration created better public water systems while waste treatment facilities increased, and sewage construction grew. These advancements protected water quality and led to a decrease in typhoid fever (Barrett & Stanberry, 2009).
Typhoid fever is most common in individuals ages 5-19 in endemic areas. Only occurring in humans, typhoid is frequently transmitted through ingestion of food or water contaminated by patients and carriers. However, transmission is also possible through direct person to person contact (Colomba et al, 2008). Symptoms of typhoid fever include fever, abdominal pain, small red spots on the abdomen and chest, severe diarrhea, and fatigue (Vyas, 2021). Illness usually lasts 4-7 days; however, it can last longer if the infection spreads to distant organs and the bloodstream (CDC, 2013).
After an individual is infected, S. typhimurium adheres to the small intestine and enters the submucosal region of the ileum, the final segment of the small bowel, before causing an increase in growth of the Peyer’s patches. The organism spreads from the Peyer’s patches through the bloodstream and lymphatic system and begins replication in the reticuloendothelial system, eventually causing symptoms that will be observed in a doctor’s office (Figure 1). After replication, organisms will stay in the macrophages of the liver, spleen, and bone marrow (Ashurst et al, 2022). Even after proper antibacterial treatment, 1-5% of typhoid patients become chronic carriers, meaning the patient excretes bacteria in their stool for over a year (Colomba et al, 2008).
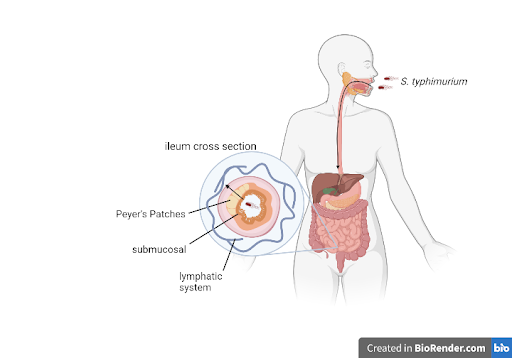
| Figure 1. Illustration of the process of S. typhimurium infecting the body through the small intestine to the lymphatic system. |
|---|
To prevent the spread of S. typhimurium in vulnerable populations, we hope to create a detector to stop individuals from ingesting contaminated food and water. Our solution would be quick, affordable, and reusable, allowing households to test the food and water they ingest for S. typhimurium and help stop the spread of disease.
Systems level
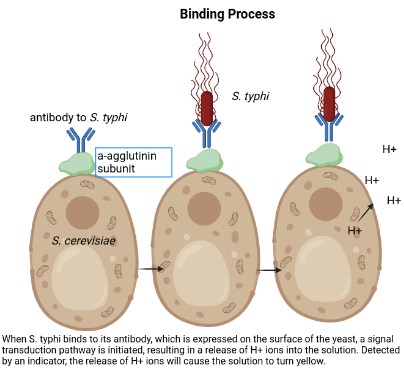
In order to detect S. typhimurium in samples of food, we will express the antibody to S. typhimurium on the surface of S. cerevisiae. The antibody will be “glued” onto the surface of the yeast with a-agglutinin complex subunits (Aga1p and Aga2p) (Figure 2) (Valldorf et al, 2021). The yeast displaying the antibody on its surface will be placed in liquid media to reproduce, and a small portion will be removed each time in order to test a sample for S. typhimurium contamination. The food sample will be placed in the portion of yeast solution for testing.
The antibody to S. typhimurium will be constructed using the Golden Gate cloning approach, which allows for multiple DNA fragments to be combined into one functional antibody. Both destination and entry vectors, along with the Type IIS enzyme and ligase, are placed in a single tube. The approach ensures, through the inclusion of Type IIS enzymes, that the selected DNA fragments combine to form the intended antibody (Valldorf et al, 2021.)
If S. typhimurium in the sample binds to the antibody, it will initiate a signal transduction pathway that remains unknown. This pathway will trigger a release of protons into the surrounding liquid media. A pH indicator, such as Bromophenol blue, will be added to the solution, and the increase in proton concentration will trigger a color change from blue to yellow. This color change will indicate S. typhimurium contamination (Figure 3).
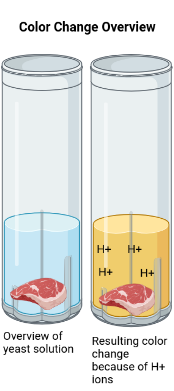
| Figure 3. Illustration of the color change that occurs as a result of a food sample contaminated with S. typhimurium being added to the yeast solution. |
|---|
Device level
We chose S. cerevisiae as a chassis because it is well-known and can self-generate. Using a-agglutinin complex subunits (Aga1p and Aga2p), we will attach the antibody to S. typhimurium, which will be constructed with the Golden Gate Cloning approach, to the yeast surface, similar to previous experiments that used this technique for yeast surface display. (Weicher et al, 2020)
In order for our solution to be successful after S. typhimurium binds to its antibody, it must initiate a signal transduction pathway that will result in a release of protons into the surrounding media (Figure 3). Though that pathway has not yet been defined, it is essential to the function of the system; thus, further research will focus on identifying this pathway. Previous research has proven that synthetic biology can be used to initiate transmembrane proton release through the capture of light energy. (Hirschi et al, 2019) Thus, we will investigate methods to initiate proton release related to both harnessing light energy and high-energy substrates.
This proton release will be detected with a pH indicator. Based on our research, we have identified Bromophenol blue as a potential candidate because it changes color at a pH slightly lower than that of yeast. (Zhao et al, 2001) This range would be beneficial because it would detect an increase in the normal acidity of S. cerevisiae. However, as we construct and test our solution, we may reconsider indicator choice based on our results.
Parts level
To construct the antibody to S. typhimurium, we plan to use the Golden Gate Cloning approach. We chose this approach because multiple fragments of DNA can be cloned simultaneously in order to produce one antibody to display on the surface of the yeast (Valldorf et al, 2021). As outlined in recent research, there are two ways to construct an antibody: a two-directional system and a bicistronic system. Within the two-directional system, the Gal1 promoter is used to express the heavy chain, and the Gal10 promoter is used to express the light chain. Within the bicistronic system, ribosomal skipping is used to create the antibody. (Rosowski et al, 2018). Based on further research, we will choose the system of antibody construction that is most relevant to the context of our research.
In order to allow the S. typhimurium antibody to bind to S. cerevisiae, we will use an a-agglutinin protein, which is expressed by S. cerevisiae and alpha cells. Agglutin works when the antigen mixes with its corresponding antibody at the correct pH and temperature, which is lower than room temperature, explaining why our yeast solution will need to be refrigerated.
The a-agglutinin is composed of two subunits: Aga1p and Aga2p. The Aga1p subunit anchors, or “glues,” a-agglutinin to the S. cerevisiae cell. The Aga2p subunit adheres to the antibody, attaching the antibody to S. cerevisiae. Both of these subunits are expressed in alpha and a cells, so we will be able to attach them to the S. cerevisiae cell. (Nobel, 1995)
The specific pathway by which proton release will be initiated has not yet been defined. In order to construct our solution, we must define this pathway; this is the focus of future research. Although we have not yet identified a specific pathway, we have concluded that a transmembrane proton pump relies either on light energy or high-energy substrates, such as ATP. In 2019, a group developed a light-energized synthetic biology proton pump, proving that such a pathway can be created using synthetic biology. (Hirschi et al, 2019) Based on this research, it is likely that a similar signaling pathway can be found and used in our yeast solution.
Safety
The system will be developed in a lab, using the Golden Gate Cloning approach to construct the antibody to S. typhimurium. Then, the antibody constructed will be “glued” onto the surface of the yeast with a-agglutinin complex subunits (1p and Aga2p). Both techniques have been previously tested with S. cerevisiae and shown to be safe, decreasing any risk of harm to developers. (Valldorf et al, 2021)
When testing the solution, live S. typhimurium cultures must be used in order to determine its functionality. Although S. typhimurium can cause serious illness if ingested, precautions will be taken in order to prevent the spread of disease. When working with S. typhimurium, which is done in a BSL-2 lab, developers will wear PPE and avoid touching their mouth, face, or other surfaces. After concluding an experiment, PPE must be disposed of, surfaces must be wiped down with antibacterial wipes, and developers must wash their hands thoroughly. Access to a sink, biological safety cabinet, and an eyewash station is required. No food will be allowed in the lab.
Because the solution is self-regenerating, no potentially harmful preservatives will be added to the solution. By avoiding the use of preservatives, which can be harmful if ingested or disposed of incorrectly, we will be able to avoid harming users and the environment.
However, because the yeast solution is self-regenerating, it has the potential to mutate and spread that mutation to later generations. Though many mutations are harmless or have little effect, there is potential for a harmful mutation to spread. After we construct the yeast solution and conduct tests to determine how prevalent significant mutations are, we will be able to evaluate the risks posed by potential mutations. If mutations appear to pose a significant threat, we can implement a testing program where users can periodically send a sample to a lab in order to ensure that their yeast solution does not contain harmful mutations. However, as of now, it does not seem to pose a significant risk; there does not seem to be a greater risk of mutations than people working with baking yeast.
Discussions
Our project is an effective solution for testing in vulnerable populations because it is self-regenerating and simple to use, making it accessible for everyday use. Because yeast solution can be grown in a home with only a small portion used per test, similar to a starter used in baking bread, it would be a one-time purchase for a home or community, minimizing cost.
Unlike conventional techniques, which are generally lab-based tests such as a PCR or ELISA, our solution can be easily used outside the lab, increasing accessibility in areas where people do not have reliable access to lab-based tests. Because the populations most vulnerable to S. typhimurium contamination also lack reliable access to lab-based screenings, they are often unable to test for contamination, rendering conventional S. typhimurium detectors useless. Although our solution would need to be refrigerated, increasing access to refrigeration is simpler than increasing access to laboratories. We are also exploring the possibility of freeze-drying the yeast, which would eliminate the need for refrigeration and increase ease of transport. Because our solution, unlike many conventional detectors, can be used outside the lab, it will allow for people in vulnerable areas to access testing, even if there is no lab nearby.
The color change as an indicator of S. typhimurium contamination would also be an easy-to-interpret result, allowing people who may be unfamiliar with more complex scientific techniques to use and understand it. Conventional techniques, which are generally lab-based tests such as a PCR or ELISA, cannot be performed in the average home. Results are technical, and people without scientific backgrounds would not be able to interpret the results even if they had access to those tests. By removing the barrier presented by complicated processes, our testing solution will be accessible to the average person, increasing testing for S. typhimurium and increasing food safety.
However, our solution does present some challenges, mainly through a lack of testing. Because we have not yet constructed the yeast, we have not yet been able to determine the intensity of the color change in proportion to the level of S. typhimurium contamination. In order to construct and test our solution, we must identify the signal transduction pathway that causes the release of protons when S. typhimurium binds to the antibody presented on the surface of the yeast. Then, after we construct the solution, we can resolve our challenge by adding varying amounts of S. typhimurium and then observing the resulting color change.
Another challenge that can be resolved after yeast construction is the media in which it will be stored. After we construct the yeast, we can determine the liquid media in which it will grow best. However, because our design depends on a pH indicator, we must ensure that the media does not change the pH significantly. After construction, we can test the pH of the media, and depending on the result, we can reconsider our choice of indicator in order to provide accurate results.
Next Steps
In order to successfully construct our solution, we must enumerate the pathway by which a release of protons can be initiated after S. typhimurium binds to its antibody. We must also determine which system of the Golden Gate Cloning approach we will use in order to construct the antibody to S. typhimurium.
In order to better understand how to create our solution, which uses yeast surface display, we plan to complete an experiment about yeast transformation. Using the Golden Bread experiment from BioBuilder, we plan to bioengineer yeast to contain more vitamins. After completing this experiment, we will better understand the process of yeast transformation, which will help us better understand how to construct a yeast surface display. Once we have completed this experiment and conducted further research on the pathway that causes proton release, we will be able to begin construction of our solution.
Author contributions
H.S, L.B., C.L., and R.R. conceived and researched the idea. H.S. took the lead in writing the manuscript, the device level, the systems level, the safety section, discussion, and next steps. R.R. wrote the background section, cited sources, and created graphics. C.L. created the graphics and wrote the parts level, and L.B. wrote the acknowledgements, cited sources, and contributed to the safety section. All authors provided critical feedback and helped shape both the research and the manuscript.
Acknowledgements
We would like to thank our club advisors Alexis Dimitriou and Sarah Hanna for their help in developing our project. We would also like to thank the BioBuilders organization for providing us with research resources and for hosting us at the final assembly, where we were able to receive feedback and incorporate it into our project. We would also like to thank our BioBuilder mentor Natalie Kuldell for all of her help, especially in wording our project and sending us resources and articles to use in our research.
References
Als, D., Radhakrishnan, A., Arora, P., Gaffey, M. F., Campisi, S., Velummailum, R., Zareef, F., & Bhutta, Z. A. (2018). Global trends in typhoidal Salmonellosis: A systematic review. The American Journal of Tropical Medicine and Hygiene, 99(3 Suppl), 10. https://doi.org/10.4269/ajtmh.18-0034
Ashurst, J. V., Truong, J., & Woodbury, B. (n.d.). Typhoid Fever (Salmonella Typhi). StatPearls Publishing. https://www.ncbi.nlm.nih.gov/books/NBK519002/
ATCC. (n.d.). Salmonella enterica subsp. enterica (ex kauffmann and edwards) le minor and popoff serovar typhimurium. https://www.atcc.org/products/14028
Barrett, A. D., & Stanberry, L. R. (2009). Vaccines for biodefense and emerging and neglected diseases. Academic Press
Cockerham, W. C. (2016). International Encyclopedia of public health. Elsevier.
de Nobel, H., Pike, J., Lipke, P. N., & Kurjan, J. (1995). Genetics of a-agglutunin function in Saccharomyces cerevisiae. Molecular and General Genetics MGG, 247, 409-415. https://doi.org/10.1007/BF00293141
Government of Canda. (n.d.). Pathogen safety data sheets: Infectious substances – Salmonella enterica spp. https://www.canada.ca/en/public-health/services/laboratory-biosafety-biosecurity/pathogen-safety-data-sheets-risk-assessment/salmonella-enterica.html
Hirschi, S., Fischer, N., Kalbermatter, D., Laskowski, P. R., Ucurum, Z., Müller, D. J., & Fotiadis, D. (2019). Design and assembly of a chemically switchable and fluorescently traceable light-driven proton pump system for bionanotechnological applications. Scientific Reports, 9(1), 1046. https://doi.org/10.1038/s41598-018-37260-9
iGEM. (n.d.). General Design of the Interleukin Yeast Biosensor. https://2020.igem.org/Team:UCopenhagen/Design
Jong, E. C., & Stevens, D. L. (Eds.). (2021). Netter’s infectious diseases Elsevier Health Sciences.
Kim, S. H., & Bansal, J. (2022). A rare case of typhoid fever in the United States associated with travel to Mexico. Cureus, 14(2). https://doi.org/10.7759/cureus.22316
Pollard, A. J. (2022). Understanding how salmonella typhi infects humans. NIH. https://clinicaltrials.gov/ct2/show/study/NCT03889067
Quah, S. R., & Cockerham, W. C. (2016). International Encyclopedia of public health (2nd ed.). Elsevier.
Rosowski, S., Becker, S., Toleikis, L., Valldorf, B., Grzeschik, J., Demir, D., Willenbücher, I., Gaa, R., Kolmar, H., Zielonka, S., & Krah, S. (2018). A novel one-step approach for the construction of yeast surface display Fab antibody libraries. Microbial Cell Factories, 17, 1-11. https://doi.org/10.1186/s12934-017-0853-z
Srisa-Art, M., Boehle, K. E., Geiss, B. J., & Henry, C. S. (2018). Highly sensitive detection of Salmonella typhimurium using a colorimetric paper-based analytical device coupled with immunomagnetic separation. Analytical Chemistry, 90(1), 1035-1043. https://doi.org/10.1021/acs.analchem.7b04628
Valldorf, B., Hinz, S. C., Russo, G., Pekar, L., Mohr, L., Klemm, J., Doerner, A., Krah, S., Hust, M., & Zielonka, S. (2021). Antibody display technologies: Selecting the cream of the crop. Biological Chemistry, 403(5-6), 455-477. https://doi.org/10.1515/hsz-2020-0377
Vyas, J. M. (2021, June 20). Typhoid fever. Penn Medicine. https://www.pennmedicine.org/for-patients-and-visitors/patient-information/conditions-treated-a-to-z/typhoid-fever
Wain, J., House, D., Zafar, A., Baker, S., Nair, S., Kidgell, C., Bhutta, Z., Dougan, G., & Hasan, R. (2005). Vi antigen expression in Salmonella enterica serovar Typhi clinical isolates from Pakistan. Journal of Clinical Microbiology, 43(3), 1158-1165. https://doi.org/10.1128/jcm.43.3.1158-1165.2005 Zhao, H., Shen, Z. M., Kahn, P. C., & Lipke, P. N. (2001). Interaction of α-agglutinin and a-agglutinin, Saccharomyces cerevisiae sexual cell adhesion molecules. Journal of Bacteriology, 183(9), 2874-2880. https://doi.org/10.1128/jb.183.9.2874-2880.2001